Filters: Tags: species distribution modeling (X)
93 results (209ms)|
Filters
Date Range
Extensions Types Contacts
Categories Tag Types
|

Average projected future (across 5 regional climate models using the A2 emissions scenario) density (birds per hectare) model of Streaked Horned Lark (Eremophila alpestris strigata) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from 1) Weather Research Forecasting Grell Model (WRFG) with boundary conditions driven by the Third Generation Coupled Global Climate Model (CGCM3); 2) Weather Research Forecasting Grell Model (WRFG) with boundary conditions driven by the Community Climate System Model (CCSM); 3) Regional Climate Model v3 (RCM3) with boundary conditions driven by the Geophysical Fluid Dynamics...

Current density (birds per hectare) model of Orange-crowned Warbler (Oreothlypis celata) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from PRISM (Daly et al. 2004) averaged for the years 1971-2000.

Current density (birds per hectare) model of Warbling Vireo (Vireo gilvus) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from PRISM (Daly et al. 2004) averaged for the years 1971-2000.
Current binomial (presence/absence) model of Brown Creeper (Certhia americana) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from PRISM (Daly et al. 2004) averaged for the years 1971-2000.
Categories: Data;
Types: Downloadable,
GeoTIFF,
Map Service,
Raster;
Tags: California,
NPLCC,
Oregon,
Washington,
birds,
Note: this data release has been superseded by version 2.0, available here: https://doi.org/10.5066/P9V54H5K We developed habitat suitability models for invasive plant species selected by Department of Interior land management agencies. We applied the modeling workflow developed in Young et al. 2020 to species not included in the original case studies. Our methodology balanced trade-offs between developing highly customized models for a few species versus fitting non-specific and generic models for numerous species. We developed a national library of environmental variables known to physiologically limit plant distributions and relied on human input based on natural history knowledge to further narrow the variable...
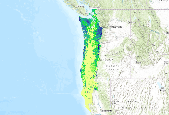
Current density (birds per hectare) model of Townsend's Warbler (Setophaga townsendi) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from PRISM (Daly et al. 2004) averaged for the years 1971-2000.

Average projected future (across 5 regional climate models using the A2 emissions scenario) density (birds per hectare) model of Rufous Hummingbird (Selasphorus rufus) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from 1) Weather Research Forecasting Grell Model (WRFG) with boundary conditions driven by the Third Generation Coupled Global Climate Model (CGCM3); 2) Weather Research Forecasting Grell Model (WRFG) with boundary conditions driven by the Community Climate System Model (CCSM); 3) Regional Climate Model v3 (RCM3) with boundary conditions driven by the Geophysical Fluid Dynamics Laboratory Global...
This dataset provides spatial predictions of habitat suitability for current (1950 – 2000 yr) and mid-Holocene (8.3 ka – 4.2 ka) intervals using hindcasting, and three separate paleo-distributions calibrated on the packrat midden archive: those without bias correction (naïve), those created with a standard method (standard), and those created with a novel alternative (modeled) incorporating a three-stage model of bias. The raster layers contained here accompany the manuscript Inman et al. 2018 and were used to evaluate utility of a novel bias correction method (modeled) over classic methods. Spatial predictions of habitat suitability were created using MaxEnt version 3.4.0 (Phillips et al., 2006), a widely-used...
We developed habitat suitability models for invasive plant species selected by Department of Interior land management agencies. We applied the modeling workflow developed in Young et al. 2020 to species not included in the original case studies. Our methodology balanced trade-offs between developing highly customized models for a few species versus fitting non-specific and generic models for numerous species. We developed a national library of environmental variables known to physiologically limit plant distributions (Engelstad et al. 2022 Table S1: https://doi.org/10.1371/journal.pone.0263056) and relied on human input based on natural history knowledge to further narrow the variable set for each species before...

Average projected future (across 5 regional climate models using the A2 emissions scenario) density (birds per hectare) model of Olive-sided Flycatcher (Contopus cooperi) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from 1) Weather Research Forecasting Grell Model (WRFG) with boundary conditions driven by the Third Generation Coupled Global Climate Model (CGCM3); 2) Weather Research Forecasting Grell Model (WRFG) with boundary conditions driven by the Community Climate System Model (CCSM); 3) Regional Climate Model v3 (RCM3) with boundary conditions driven by the Geophysical Fluid Dynamics Laboratory...

Current density (birds per hectare) model of Willow Flycatcher (Empidonax traillii) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from PRISM (Daly et al. 2004) averaged for the years 1971-2000.

Current density (birds per hectare) model of Savannah Sparrow (Passerculus sandwichensis) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from PRISM (Daly et al. 2004) averaged for the years 1971-2000.
The "_archive_workflow_FinalModel.zip" data bundle is comprised of the metadata and Vistrails workflow that contains the following history nodes, which contain modeling workflows: "NLCD2016 state bckgrnd" and "NLCD2016 wostate bckgrnd". These nodes produced the following 9 output rasters: 1) Probability map (without state) 2) MPP threshold (without state) 3) Five percent threshold (without state) 4) Ten percent threshold (without state) 5) Probability map (with state) 6) MPP threshold (with state) 7) Five percent threshold (with state) 8) Ten percent threshold (with state) 9) Maxent MESS map
The 'archive_raster_inputs.zip' data bundle contains '_archive_raster_inputs_XX.tif' and archive_raster_inputs_XX.xml where XX is the name of 1 of 8 input rasters that were created and used to generate these model results. The original layers and sources used to produce each predictor, as well as processing steps, are specified in each .xml file.
We developed habitat suitability models for three invasive plant species: stiltgrass (Microstegium vimineum), sericea lespedeza (Lespedeza cuneata), and privet (Ligustrum sinense). We applied the modeling workflow developed in Young et al. 2020, developing similar models for occurrence data, but also models trained using species locations with percent cover ≥10%, ≥25%, and ≥50%. We chose predictors from a national library of environmental variables known to physiologically limit plant distributions (Engelstad et al. 2022 Table S1) and relied on human input based on natural history knowledge to further narrow the variable set for each species before developing habitat suitability models. We developed models using...
We developed habitat suitability models for occurrence of three invasive riparian woody plant taxa of concern to Department of Interior land management agencies, as well as for three dominant native riparian woody taxa. Study taxa were non-native tamarisk (saltcedar; Tamarix ramosissima, Tamarix chinensis), Russian olive (Elaeagnus angustifolia) and Siberian elm (Ulmus pumila) and native plains/Fremont cottonwood (Populus deltoides ssp. monilifera and ssp. wislizenii, Populus fremontii), narrowleaf cottonwood (Populus angustifolia), and black cottonwood (Populus balsamifera ssp. trichocarpa and ssp. balsamifera). We generally followed the modeling workflow developed in Young et al. 2020. We developed models using...
Categories: Data;
Tags: Elaeagnus angustifolia,
Populus spp.,
SAHM,
Software for Assisted Habitat Modeling,
Species Distribution modeling,
This dataset contains predictions of habitat suitability of reed canarygrass (Phalaris arundinacea) in Upper Mississippi River floodplain forest understories from Pool 3 to Pool 13. Predictions were created using three machine learning algorithms (Bayesian additive regression trees, boosted trees, and random forest). This dataset contains rasters that provide habitat suitability predictions for each 12m raster cell that had forested landcover in 2010. In addition to one raster for each of the three algorithms an ensemble (mean prediction of all three algorithms) prediction raster for each pool is provided. The presence/absence observations used to train the model are contained in a .csv file with each plot location....
This project used species distribution modeling, population genetics, and geospatial analysis of historical vs. modern vertebrate populations to identify climate change refugia and population connectivity across the Sierra Nevada. It is hypothesized that climate change refugia will increase persistence and stability of populations and, as a result, maintain higher genetic diversity. This work helps managers assess the need to include connectivity and refugia in climate change adaptation strategies. Results help Sierra Nevada land managers allocate limited resources, aid future scenario assessment at landscape scales, and develop a performance measure for assessing resilience.
Categories: Data,
Project;
Tags: 2011,
2013,
CA,
California Landscape Conservation Cooperative,
Conservation Design,
Future density (birds per hectare) model of Brown Creeper (Certhia americana) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from the Canadian Regional Climate Model (CRCM) with boundary conditions driven by the Community Climate System Model (CCSM) averaged for the years 2041-2070 and available from http://www.narccap.ucar.edu/.
Categories: Data;
Types: Downloadable,
GeoTIFF,
Map Service,
Raster;
Tags: California,
Oregon,
Washington,
birds,
boosted regression tree model,
Current probability of occurrence model of Scrub Jay (Aphelocoma californica) using a Boosted Regression Tree model (Hastie & Tibshirani 2000) informed by breeding season avian point count data, modeled vegetation types, and climate data from PRISM (Daly et al. 2004) averaged for the years 1971-2000.
Categories: Data;
Types: Downloadable,
GeoTIFF,
Map Service,
Raster;
Tags: California,
California,
NPLCC,
Oregon,
Oregon,
|

|